My Projects
Molecular Docking of HIV Biomarkers with Drug and Peptide-based Treatments
This project uses computer simulations to study how different drugs interact with HIV-1 protease, a key enzyme that helps the virus replicate. By modeling how these drugs bind to the enzyme, the research aims to identify which ones might be most effective in stopping the virus, including new drugs that could work against drug-resistant forms of HIV.
Paper


COVID-19 Severity Prediction via Stacking Ensemble ML Approach
This awarded research addresses the challenge of using diverse data types - clinical, proteomic, and metabolomic - to predict COVID-19 case severity. By implementing a stacking ensemble approach with each of the three ML models representing a different data type, this offers a more comprehensive perspective on COVID-19 severity prediction.
Github
Alzheimer's Disease Neurological Change (ADNC) Prediction
Leveraged the Seattle Alzheimer's Disease Brain Cell Atlas (SEA-AD) data, including neuropathological data, to predict ADNC severity with machine learning. Interpretability of ML model was visualized using LIME and SHAP techniques.
Github


Association between Kidney Disease and Depression
Investigating the relationship between kidney disease and depression in the US population, using data from the Behavioral Risk Factor Surveillance System (BRFSS) 2021. Univariate and multivariate logistic regression analyses, considering a complex survey design, were conducted to explore this association.
Github
Computational Drug Discovery
Built machine learning models using ChEMBL bioactivity data with a target protein, beta-secretase 1, to make predictions on pIC50 and data-driven insights useful for drug discovery.
Github

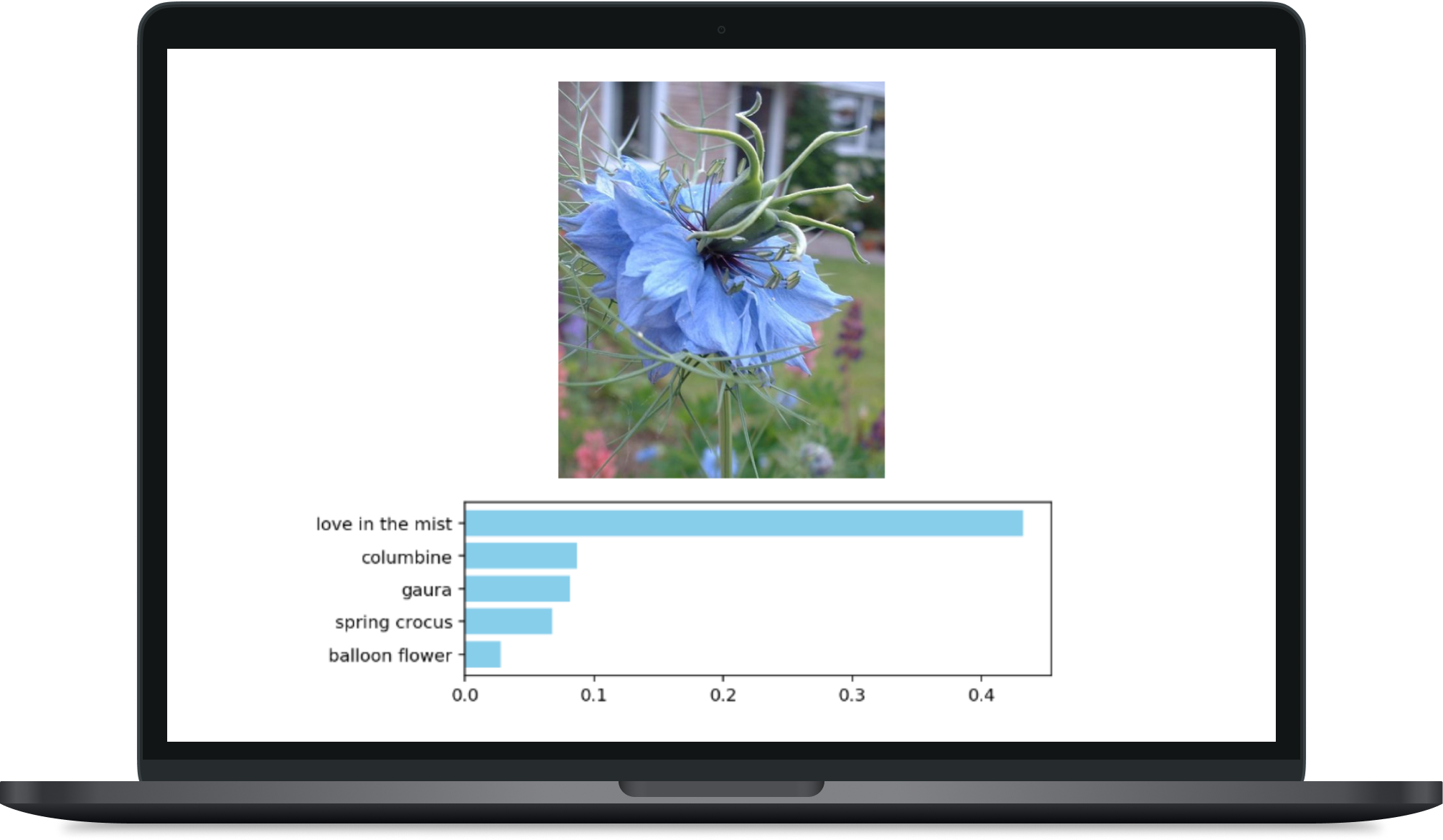
Flower Image Classifier
A custom image classifier built with PyTorch. It utilizes a VGG-16 CNN architecture to train a model on a flower image dataset. The trained classifier predicts flower type based on input images.
Github
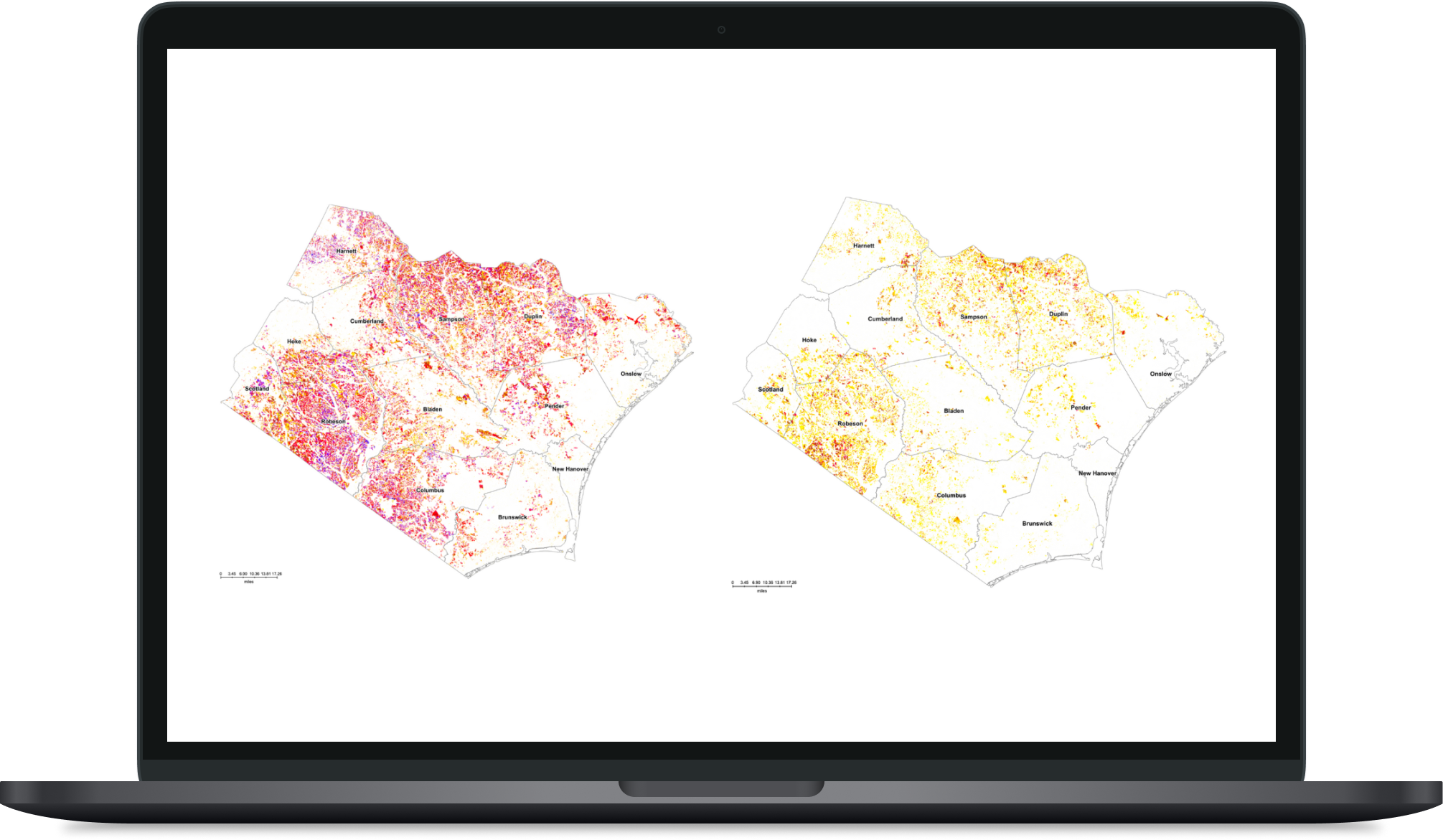
Double Cropped Acreage Variation in Southeastern NC
A research project focused on analyzing USDA cropland data in southeastern NC and fitting econometric models based on the relationship between double-cropped acreage and commodity prices.
Poster

Dog Breed Image Classifier
A pre-trained dog breed image classifier that identifies dog images and dog breeds. It also includes functions to determine which CNN architectures work best for specific criteria.
Github